CluSeek
ClusterSeeker
What is CluSeek?
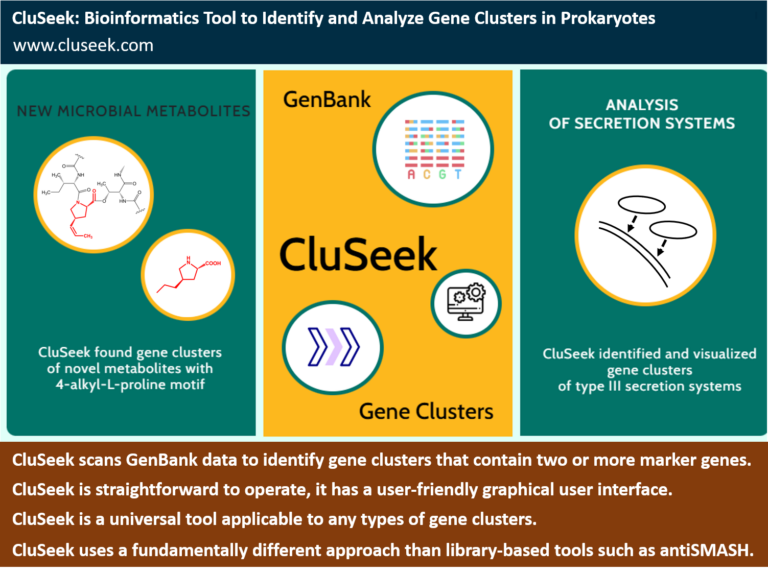
CluSeek is a versatile tool for identifying gene clusters in GenBank data. It first searches for homologs of two or more user-selected marker genes and identifies regions where these homologs are colocalized within bacterial genomes. The resulting sequences, including the markers and their neighboring genes, are then visualized as gene clusters. These visualized clusters can be further explored and analyzed using CluSeek’s built-in functions through a user-friendly interface.
CluSeek can be applied to any type of gene cluster, regardless of the encoded phenotype, including clusters involved in the biosynthesis of specialized metabolites, secretion systems, complex metabolic pathways, virulence factors, chemotactic motility, and more.
CluSeek enables efficient genome mining of GenBank data using a strategy distinct from mainstream tools such as antiSMASH, which is widely used for specialized metabolite analysis.
Research of specialized Metabolites:
Is CluSeek similar to antiSMASH?
Genome mining of biosynthetic gene clusters is typically performed with tools like antiSMASH. AntiSMASH identifies clusters within individual genomes using predefined rules derived from well-characterized biosynthetic classes. In contrast, CluSeek operates on a fundamentally different principle: it scans the entire GenBank without relying on prior rules, internal libraries, or knowledge of known clusters. In fact, gene clusters identified by CluSeek can subsequently be exported for their analysis with antiSMASH, bridging the two approaches.
Is CluSeek easy to use?
Yes. CluSeek follows a simple download-and-use principle and has a user-friendly graphical interface, making it accessible to users without IT expertise. To help users navigate the interface, we have also prepared step-by-step video tutorials available on YouTube.
Are there any limitations to using CluSeek?
CluSeek was developed for prokaryotic genomes; however, preliminary testing shows that it can also handle sequencing data from eukaryotes. It is fully compatible with Windows and macOS, and is also accessible on Linux via a Python package.
Can I set up CluSeek now?
Set Up:
- Download CluSeek for Windows here , and for macOS here
- and extract the .zip file
- Windows only: If you are setting-up CluSeek on Windows for the first time, you will need to install the Windows redistributables (vc_redist.x86) included with CluSeek, or download and install the latest version from microsoft.
- That’s it. Double click the CluSeek.exe application in the folder and start new analysis.
Note that CluSeek is also available as a Python package named cluseek on the Python Package Index for Windows, macOS and Linux.
If you have python installed, you can install the package using the command:
pip install cluseek
and run it using the comand:
python -m cluseek
conda create --platform osx-64 --name cluseek_env python=3.9
conda activate cluseek_env
pip install cluseek
python -m cluseek
You can find our source code on GitHub.
Video tutorials are available on our YouTube channel.
CluSeek v2.0.2
Minor changes:
– Hidden the abort button in the download dialog until abort functionality can be re-worked
– Hidden unused radio button “Other” in the protein group network view
Bug fixes:
– Updated FASTA protein export user-specified identifiers for clusters, not the internal numerical ones
– Fixed offline mode button
– Included coding sequence strand information in the GenBank format export
Cosmetic changes:
– Adjusted UI alignment in the colocalization histogram
– Changed spelling of E-value in colocalization histogram
– Changed capitalization for drop down menus on the toolbar
– Corrected typo in the Cluster view screen
– Rewrote some error messages to improve clarity
You can send us your feedback or suggestions at cluseek@biomed.cas.cz
Should you require an older version of CluSeek, please do not hesitate to contact us via email.
Institue of Microbiology
Institute of Microbiology, CAS
Vídeňská 1083, 142 20 Prague 4 - Krč
The Czech Republic
BIOCEV
Institute of Microbiology, CAS
Průmyslová 595, 252 50 Vestec
The Czech Republic
Contact Us
lab111@biomed.cas.cz
+420 241 062 371
